A quality control test on Salmonella typing has revealed positive results among European laboratories.
National Reference Laboratories (NRLs) of the 28 European Union member states were able to assign the correct name to 98 percent of the strains tested.
The objective of the test in November 2017 was to evaluate whether the typing of Salmonella strains by the NRLs in the EU were carried out uniformly and if results were comparable.
The inter-laboratory comparison study is organized by the European Union Reference Laboratory (EURL) for Salmonella at the National Institute for Public Health and the Environment (RIVM) in the Netherlands.
Salmonella strains as part of the test included Agona, Weltevreden, Typhimurium, Stanley, Enteritidis and Infantis. Just less than half of the 35 participating labs had a serotyping frequency of daily in 2017 with only one lab doing such work monthly.
The standard method for typing Salmonella is serotyping but 15 labs performed typing at DNA level using Pulsed Field Gel Electrophoresis (PFGE). This more detailed method can be used to trace the source of a contamination. For quality control, participants received another 11 strains of Salmonella for PFGE.
Overall, 90 percent of the scores were good or excellent. However, four of the 15 images resulted in a poor score on at least one of the seven parameters. These four images were unsuitable for inter-laboratory database comparison of these PFGE profiles.
Ten also processed a common gel in the software BioNumerics and all of them were able to analyze the PFGE profiles in this computer program.
Performance of EU NRLs in Salmonella typing is assessed annually by testing ability to identify 20 strains. NRLs from countries outside the EU participate on a voluntary basis. The Former Yugoslav Republic of Macedonia and Serbia as well as Iceland, Norway, Switzerland and Israel took part in the 2017 assessment.
All 35 labs performed serotyping. Twenty obligatory Salmonella strains plus one optional strain were selected by the EURL-Salmonella for serotyping. They had to be typed according to the method routinely used in each lab, following the White-Kauffmann-Le Minor scheme.
The O-antigens were typed correctly by 31 of 35 participants which corresponded to 99 percent of the total number of strains. The H-antigens were typed accurately by 28 participants (98 percent of the strains). As a result, 28 participants also gave the correct serovar names (98 percent of all strains evaluated).
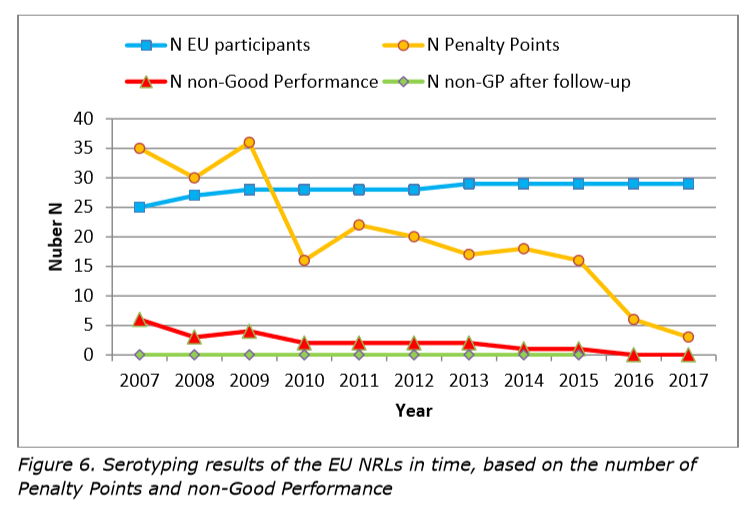
All 35 participants achieved good results in the first stage of the study based on criteria for good performance defined in 2007. Of the participants, 32 were accredited for serotyping Salmonella, mainly according to ISO 17025. The other three labs were working on such accreditation.
The number of EU NRLs with a non-good performance is low: two in the period 2010 – 2013, only one in the 2014 and 2015 studies, and none in the 2016 and 2017 tests.
RIVM also recently published results for the Salmonella typing in 2016 and 2015.
In 2016, nearly 100 percent of the strains were typed correctly for the O-antigens, 99 percent were typed accurately for the H-antigens and 99 percent were correctly named by the participants. Thirteen of 15 participating labs produced a PFGE gel of sufficient quality.
Two non-EU-NRLs did not achieve the set level of good performance and were offered a follow-up study, typing an additional 10 strains. One non-EU-NRL participated but it was not able to improve.
In 2015, 99 percent of the strains were correctly typed for the O-antigens, 97 percent were correctly typed for the H-antigens and 97 percent were correctly named.
Thirty three participants achieved good results. The lab that did not achieve this level of performance participated in a follow-up study including 10 additional strains for serotyping. This EU-NRL got good scores in the obligatory follow-up study.
(To sign up for a free subscription to Food Safety News, click here.)









