Two-thirds of 30 European countries were routinely using whole genome sequencing in 2017 for national surveillance of at least one human pathogen, according to a survey.
This figure is up from 16 countries in 2016 and none in 2013.
A European Centre for Disease Prevention and Control (ECDC) report found 29 of the 30 countries surveyed said they are either using or planning to start genomic surveillance by 2019. The only exception was Cyprus.
The agency said with a harmonization of standards, this will enable the exchange of WGS-derived data across the continent as prioritized in its roadmap for molecular surveillance and improve disease control and prevention.
The survey found the majority of national public health reference laboratories in the European Union (EU) and European Economic Area (EEA) countries have access to WGS-based typing of diverse microbial pathogens for investigations of infection, virulence, and drug resistance transmission.
Results will be used in the revision of ECDC’s roadmap on disease priorities for molecular typing integration into surveillance and epidemic preparedness.
Whole genome sequencing (WGS) with epidemiological and environmental investigations provides higher resolution and accuracy than traditional molecular typing methods such as Pulsed-Field Gel Electrophoresis (PFGE) and Multi-Locus Variable-Number Tandem Repeat Analysis (MLVA).
The ECDC said it would help to consolidate and harmonize WGS-based typing methods to enable pan-EU genomic surveillance and multi-country outbreak investigations by developing sequence and epidemiological data exchange, management and analysis systems.
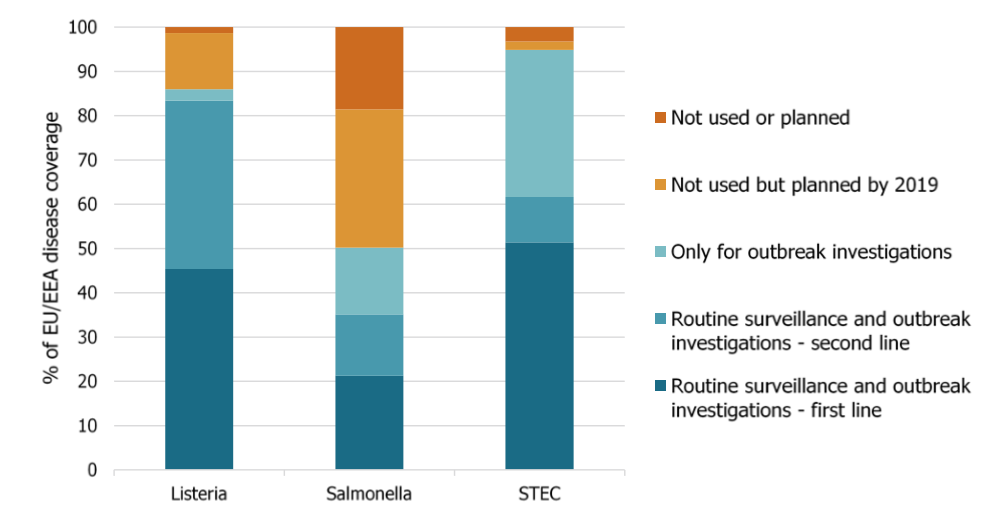
Listeria monocytogenes was the foodborne pathogen most frequently typed using WGS for surveillance and outbreak investigations last year followed by Shiga toxin producing E. coli (STEC) and Salmonella enterica.
The pathogen most comprehensively characterized by WGS as first-line typing method was Listeria monocytogenes in nine countries, followed by STEC in eight and Salmonella enterica in five.
Illumina was the most frequently used next-generation sequencing (NGS) technology for outbreak investigations or routine surveillance followed by Ion Torrent.
For WGS data storage, the public health labs in the majority of EU/EEA countries deposited raw sequence (fastq files) data in dedicated closed databases (either national or international) rather than public repositories such as ENA.
“The most frequently reported reason for this practice was the priority given to this information for national reporting and risk assessment, followed by the need for permission for scientific publication of original data. The third most cited reason was the need for personal data protection,” according to the report.
The survey found current bottlenecks mainly relate to access to bioinformatics expertise for routine WGS data analysis and access to user-friendly international nomenclature. Challenges with costs and lack of expertise still limit use by public health labs.
Core genome multi-locus allelic profiling was the most frequently used analytical approach to genotyping bacterial genomes, suggesting there are potential bioinformatics pipeline and nomenclature compatibility, said ECDC.
Among diverse bioinformatic pipelines used by the national public health reference labs for WGS data analysis, the core genome Multi-Locus Sequence Typing (cgMLST), often used with single nucleotide polymorphisms (SNP) analysis, was the most commonly used approach across pathogens.
(To sign up for a free subscription to Food Safety News, click here.)








