Pulsed-Field Gel Electrophoresis (PFGE) PFGE was developed in 1984 and has since become the gold standard for bacterial subtyping. PFGE is the primary subtyping method used by PulseNet, a network of public health laboratories that perform PFGE on foodborne disease organisms such as Salmonella, E. coli O157:H7, Shigella (11), Listeria monocytogenes (7), and Campylobacter jejuni (12). PulseNet has been responsible for detecting many foodborne disease outbreaks (3, 4, 13), and has been extremely successful. In 1999, PulseNet won the Innovations in American Government Program and in 2002, PulseNet was recognized as one of the fifteen most significant government initiative programs to have won the Innovations in American Government Award.
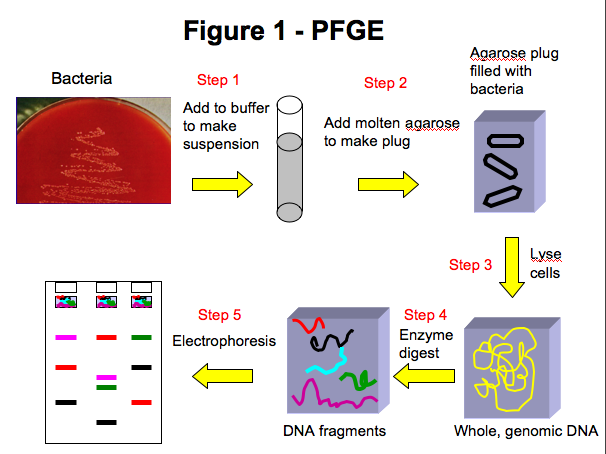
The first step of PFGE is to make a suspension of bacteria and buffer (Figure 1-step 1). Molten agarose is added to the bacterial suspension and the agarose/suspension mixture is added to a mold (Figure 1-step 2). The agarose is allowed to solidify then added to a lysis solution which breaks open the bacteria releasing the DNA. Subsequent washing steps remove extracellular debris leaving intact, circular genomic DNA in the agarose plug (Figure 1-step 3). A small piece of the plug is cut off and added to a restriction endonuclease mixture which cleaves DNA at a specific sequence (Figure 1-step 4) resulting in 10-20 DNA fragments of varying sizes. The large DNA fragments are then separated by size by pulsed-field gel electrophoresis (PFGE) (Figure 1-step 5). PFGE facilitates the migration of the large DNA fragments through the agarose gel by regularly changing the direction of the electrical field during electrophoresis, allowing the fragments to maneuver through the agarose. The smaller DNA fragments will move faster through the agarose than the larger fragments and the result is a pattern of DNA fragments (also called bands). The pattern from one isolate can be compared to other patterns to determine whether the samples may have originated from a common source. PFGE Advantages -Easily applied to different species -Patterns consistent within and between laboratories (strict adherence to standard conditions is necessary) -PFGE generally yields a high amount of pattern diversity PFGE Drawbacks -Labor-intensive -Relatively slow (approximately 24 hours to completion) -Complex patterns challenging for inter-laboratory pattern comparisons -One mutation can yield differences in several fragments (PFGE cannot determine phylogenic relationships) -Dependant on isolated bacteria Multiple-Locus Variable Number Tandem Repeat Analysis (MLVA) Multiple-locus variable number tandem repeat analysis (MLVA) is a subtyping method that utilizes short, repetitive regions of DNA called variable-number tandem repeats (VNTR) which may be present in different numbers of repeats between strains. These variable number tandem-repeat (VNTR) regions have been identified in many species of bacteria. Multiple VNTR regions can be analyzed in one typing method (MLVA) to differentiate bacteria within a species. Most VNTR regions used in MLVA are small, containing between 3 and 21 base pair repeats.
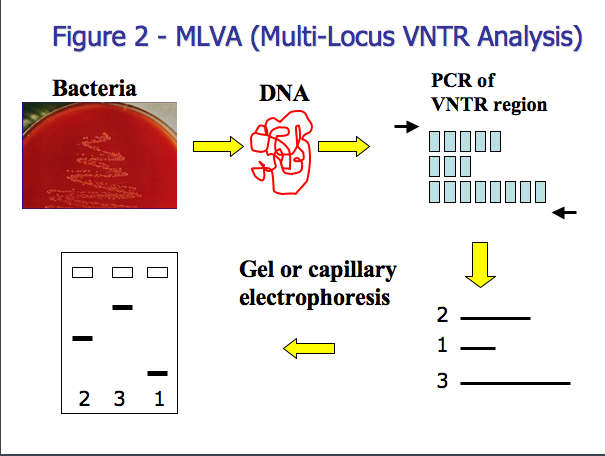
There are several steps involved in MLVA (Figure 2). The first step involves amplifying the VNTR using polymerase chain reaction (PCR is a common method used in molecular biology to amplify a specific region). PCR is used to amplify the conserved DNA outside the VNTR in addition to the variable VNTR region. The fragment size is dependant on the number of repeats for a sample. The size of the resulting fragment is determined by either horizontal or capillary electrophoresis (2). Horizontal electrophoresis uses traditional agarose gels to estimate the size of DNA fragments while capillary electrophoresis uses a DNA sequencer for fragment size analysis. Capillary electrophoresis is a particularly attractive option because you can attain extremely precise measurement of the fragments’ size (often within one base pair), which is ideal for large databases and inter-laboratory comparison.
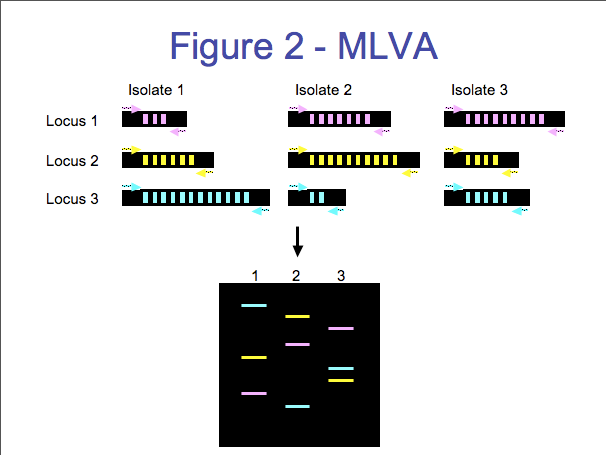
Detection of candidate loci for MLVA is dependant on knowledge of the genome for a given species. Due to the explosion of bacterial genomes that have been fully sequenced over the last decade, MLVA is becoming a more common subtyping method. MLVA has been successfully used for many genera of bacteria, including E. coli O157:H7 (8), Salmonella Typhimurium (9), Salmonella Enteritidis, and Listeria monocytogenes (many references). MLVA is used by PulseNet to further differentiate groups of E. coli O157:H7 and S. Typhimurium isolates that have the same PFGE type. MLVA Advantages -Rapid (results completed in about 5 hours) -Relatively inexpensive to perform -Portable. Amenable to inter-laboratory comparisons -Not labor intensive -Amenable for high-throughput -Loci can be added or removed to adjust diversity MLVA Drawbacks -Significant development time to create a new MLVA system -Need sequences to identify possible VNTR candidates -Sequencer is expensive and not available to many laboratories (sequencers are needed for creation of large databases) Nucleic Acid Sequence Subtyping
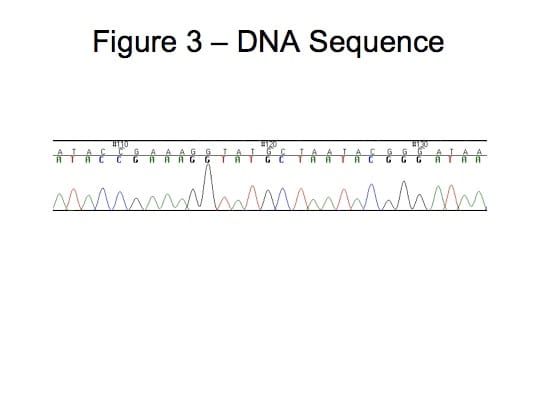
Bacteria and viruses can be differentiated within species by determining the sequence of nucleotides bases (adenine, cytosine, guanine, or thymine) for one or more loci of an organism (Figure 3). Sequencing of a single, diverse locus is necessary to create adequate discrimination for some organisms. Single locus sequence subtyping is used for genotyping norovirus (15) and hepatitis C (14). Combined sequence of multiple regions increase subtype diversity and may yield more information about the evolutionary history of a group of strains. The most common example of subtyping an organism by sequencing multiple regions is multiple locus sequence typing (MLST). MLST is the sequencing of multiple housekeeping genes (genes necessary for organism survival) for an organism. MLST methods have been created for numerous organisms, including Streptococcus pneumonia (6), Staphylococcus aureas (10), Campylobacter jejuni (5), and Neisseria meningitides (1). Housekeeping regions generally have limited diversity so techniques have been developed to sequence the more genetically diverse virulence genes, a method called multiple virulence locus sequence typing (MVLST). MVLST has been used for subtyping Listeria monocytogenes among other organisms. Sequence Subtyping Advantages -Can be used for bacteria or virus and fungus -Portable – information can be readily shared between laboratories -Can be high throughput -Can be automated -Can be used to investigate evolutionary relationships Sequence Drawbacks -Relatively expensive -Need sequence information for organism to sequence -Lacks the discriminatory power of differentiate some species (MLST) -Labor intensive if not automated -Most current methods are slow Single nucleotide polymorphisms (SNP, pronounced snip) are mutations that occur at a single nucleotide in the genome. The fifth nucleotide in the following sequences (ATAGCA, ATAGGA) is an example of a SNP. Most SNPs have only two alleles. Identification of SNPs is dependent on having the sequence of multiple organisms within a species. Analysis of multiple SNPs for an organism has recently been utilized to differentiate bacteria. The combination of multiple SNPs can be compared and used to determine the evolutionary relationships between organisms. Since such little sequence is needed to identify the SNPs, it is a method that is more amenable to rapid, high-through-put sequence methods such as pyrosequencing. SNP Advantages -Can be used for bacteria or virus and fungus -Portable-Information can be readily shared between laboratories -Can be high throughput -Can be automated -Can be used to investigate evolutionary relationships -Only small amount of sequence needed, amenable to more rapid sequence methods SNP Drawbacks -Need sequence data to identify SNPs -Many SNPs needed to increase diversity -Need automated sequencer which may not be available in many laboratories -More diversity may be needed
Dictionary of Terms- Agarose-a substance obtained from agar which is used as a medium for electrophoresis procedures
- Genome-an organisms genetic material
- Restriction endonuclease-an enzyme the cleaves DNA at a specific site to produce fragments
- Electrophoresis-a technique to migrate proteins or nucleic acid through a medium
- Locus-the chromosomal position of a gene
- Bennett DE, Cafferkey MT. Multilocus restriction typing: a tool for Neisseria meningitidis strain discrimination. J Med Microbiol. 2003 Sep;52(9):781-7.
- Boxrud D, Pederson-Gulrud K, Wotton J, Medus C, Lyszkowicz E, Besser J, Bartkus JM. Comparison of multiple-locus variable-number tandem repeat analysis, pulsed-field gel electrophoresis, and phage typing for subtype analysis of Salmonella enterica serotype Enteritidis.J Clin Microbiol. 2007 Feb;45(2):536-43.
- Centers for Disease Control and Prevention. MMWR Morb Mortal Wkly Rep. 2008 Nov 28;57(47):1277-80. Multistate outbreak of Salmonella infections associated with frozen pot pies–United States, 2007.
- Centers for Disease Control and Prevention. MMWR Morb Mortal Wkly Rep. 2009 Feb 6;58(4):85-90. Multistate outbreak of Salmonella infections associated with peanut butter and peanut butter-containing products–United States, 2008-2009.
- Dingle KE, Colles FM, Falush D, Maiden MC. Sequence typing and comparison of population biology of Campylobacter coli and Campylobacter jejuni. J Clin Microbiol. 2005 Jan;43(1):340-7.
- Enright MC, Spratt BG. A multilocus sequence typing scheme for Streptococcus pneumoniae: identification of clones associated with serious invasive disease. Microbiology. 1998 Nov;144 ( Pt 11):3049-60.
- Graves LM, Swaminathan B. PulseNet standardized protocol for subtyping Listeria monocytogenes by macrorestriction and pulsed-field gel electrophoresis. Int J Food Microbiol. 2001 Apr 11;65(1-2):55-62.
- Hyytiä-Trees E, Smole SC, Fields PA, Swaminathan B, Ribot EM. Second generation subtyping: a proposed PulseNet protocol for multiple-locus variable-number tandem repeat analysis of Shiga toxin-producing Escherichia coli O157 (STEC O157). Foodborne Pathog Dis. 2006 Spring;3(1):118-31.
- Lindstedt BA, Vardund T, Aas L, Kapperud G. Multiple-locus variable-number tandem-repeats analysis of Salmonella enterica subsp. enterica serovar Typhimurium using PCR multiplexing and multicolor capillary electrophoresis. J Microbiol Methods. 2004 Nov;59(2):163-72.
- Melles DC, van Leeuwen WB, Snijders SV, Horst-Kreft D, Peeters JK, Verbrugh HA, van Belkum A. Comparison of multilocus sequence typing (MLST), pulsed-field gel electrophoresis (PFGE), and amplified fragment length polymorphism (AFLP) for genetic typing of Staphylococcus aureus. J Microbiol Methods. 2007 May;69(2):371-5.
- Ribot EM, Fair MA, Gautom R, Cameron DN, Hunter SB, Swaminathan B, Barrett TJ. Standardization of pulsed-field gel electrophoresis protocols for the subtyping of Escherichia coli O157:H7, Salmonella, and Shigella for PulseNet. Foodborne Pathog Dis. 2006 Spring;3(1):59-67.
- Ribot EM, Fitzgerald C, Kubota K, Swaminathan B, Barrett TJ.Rapid pulsed-field gel electrophoresis protocol for subtyping of Campylobacter jejuni.J Clin Microbiol. 2001 May;39(5):1889-94.
- Wendel AM, Johnson DH, Sharapov U, Grant J, Archer JR, Monson T, Koschmann C, Davis JP. Multistate outbreak of Escherichia coli O157:H7 infection associated with consumption of packaged spinach, August-September 2006: the Wisconsin investigation. Clin Infect Dis. 2009 Apr 15;48(8):1079-86.
- Laperche S, Saune K, De´ny P, Duverlie G, Alain S, Chaix M, Gaudy C, Lunel F, Pawlotsky J, Payan C, Pozzetto B, Tamalet C, Thibault V, Vallet S, Bouchardeau F, Izopet J, Lefre`re J. Unique NS5b Hepatitis C Virus Gene Sequence Consensus Database Is Essential for Standardization of Genotype Determinations in Multicenter Epidemiological Studies. J Clin Microbiol. 2006, Feb;44: 614-616.
- Vinjé J, Hamidjaja RA, Sobsey MD. Development and application of a capsid VP1 (region D) based reverse transcription PCR assay for genotyping of genogroup I and II noroviruses. J Virol Methods. 2004 Mar 15;116(2):109-17



